News
May 2024
Analysis of how COVID-19 variants evolved to respond differently to climate is released
February 2024
Briefing note on Biodiversity and ecosystem services is released!
November 2024
Alex, MRes student in the lab, publishes his thesis on AI analysis of videos to detect bird behaviour
August 2024
Joel, MSc student in the lab, publishes his thesis on monitoring sharks with drones
May 2023
What can we learn from 100 years of phenology data from South Korea?
February 2023
EDGE 2.0 update is now out! A major update to the way a major conservation program operates co-led by our lab!
November 2022
Michael finishes the largest analysis of species’ phenology to date: >1000 species and >10,000 time-series
May 2022
The automated data pipeline that powered our COVID modelling - AREAData - is now released
February 2022
Undergraduate-led collaboration between engineering and biology reveals how to use citizen science data to assess UK rivers
November 2021
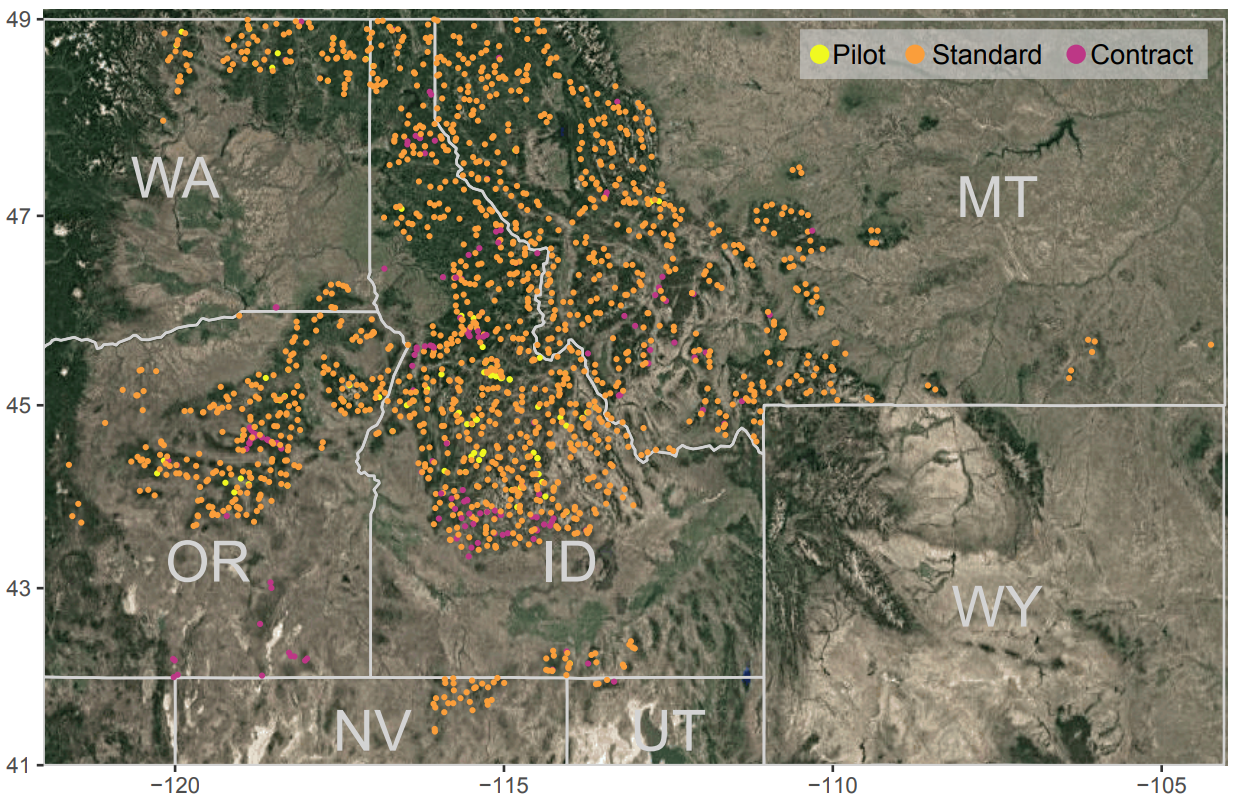
Katie’s work helping the US Forest Service prioritise restoration is released
September 2021
Tom’s manuscript on the role of environment and non-pharmaceutical interventions in COVID-19 transmission published in PNAS
May 2021
Amanda’s synthesis piece on phylogenetic modelling is Editor’s Choice in Oikos!
February 2021
Elizabeth’s first PhD chapter on fractal sampling is out!
Septmber 2020
Herbivores are at the highest risk of extinction, not carnivores: student-led Science Advances paper
June 2020
We’ve received funding from UKRI/NERC to help improve forecasting of COVID-19 and SARS-CoV-2 in the coming winter. Excited to welcome Tom Smith to the lab to work on the project!
February 2020
Excited to say the lab is moving to the Department of Life Sciences at Imperial College London!
September 2019
After almost 5 days of solid bee-catching spread over the field season, Michael is back from RMBL. Michael is exhausted, the bees are likely relieved :p 
August 2019
The Pearse Lab (Amanda, Austin, Elizabeth, Jake, and Katie) went to the Ecological Society of America meeting to present!
July 2019
Katie’s manuscript on using phylogenetic diversity metrics to efficiently do real-world conservation prioritization is submitted! Read the pre-print